Build a RAG App with Streamlit and Snowflake Cortex
Overview
Retrieval-Augmented Generation (RAG) enhances LLM responses by grounding them in your own data. Instead of relying solely on a model's training knowledge, RAG retrieves relevant documents and uses them as context thereby reducing hallucinations and enabling accurate, domain-specific answers.
In this quickstart, you'll build a complete RAG pipeline using Snowflake Cortex Search. Starting from raw documents, you'll learn data preparation, text chunking, creating a search service, building a conversational RAG chatbot, and evaluating quality with TruLens.
This tutorial distills key lessons from the #30DaysOfAI learning challenge into a hands-on guide.
What You'll Learn
- How to prepare and chunk documents for RAG (Days 16-17)
- How to generate embeddings for semantic search (Day 18)
- How to create and configure a Cortex Search service (Day 19)
- How to query Cortex Search for semantic retrieval (Day 20)
- How to build a RAG chatbot with grounded responses (Day 21)
- How to add guardrails to keep responses on-topic (Day 22)
- How to evaluate RAG quality with TruLens and AI Observability (Day 23)
What You'll Build
A complete RAG system including a Cortex Search service for customer reviews and a conversational chatbot that answers questions grounded in retrieved documents, with TruLens evaluation for quality metrics.
Prerequisites
- Access to a Snowflake account
- Basic knowledge of Python and Streamlit
- A Snowflake warehouse with Cortex Search and LLM access
Getting Started
Clone or download the code from the 30daysofai GitHub repository:
git clone https://github.com/streamlit/30DaysOfAI.git cd 30DaysOfAI/app
The app code for this quickstart:
Requirements
Save the appropriate requirements content to requirements.txt in your project directory, or add them to your Streamlit in Snowflake app's package configuration.
Days 16-22 share the same core dependencies:
streamlit snowflake-snowpark-python>=1.18.0,<2.0 snowflake>=1.0.0 pypdf pandas
Day 23 (TruLens Evaluation) requires additional packages:
streamlit snowflake-snowpark-python>=1.18.0,<2.0 snowflake>=1.0.0 pypdf pandas trulens-core trulens-providers-cortex trulens-connectors-snowflake>=2.5.0 setuptools<81
Extract and Upload Documents
The foundation of any RAG system is well-prepared data. In this section, you'll upload documents, extract their text content, and save to Snowflake.
Note: This section is repurposed from Day 16 of the #30DaysOfAI learning challenge.
Snowflake Connection
The app establishes a Snowflake connection that works both in Streamlit in Snowflake (SiS) and locally. In SiS, it uses the active session; locally, it reads credentials from st.secrets:
try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create()
Database Configuration
Users configure the target database, schema, and table name through text inputs. Session state persists these values across reruns:
if 'database' not in st.session_state: st.session_state.database = "RAG_DB" if 'schema' not in st.session_state: st.session_state.schema = "RAG_SCHEMA" if 'table_name' not in st.session_state: st.session_state.table_name = "EXTRACTED_DOCUMENTS"
File Upload and Text Extraction
The file uploader accepts TXT, Markdown, and PDF files. For PDFs, pypdf.PdfReader extracts text from each page. A progress bar shows extraction status for batch uploads:
uploaded_files = st.file_uploader( "Choose file(s)", type=["txt", "md", "pdf"], accept_multiple_files=True )
Replace vs Append Mode
A checkbox lets users choose between replacing existing data or appending to it. The default is based on whether the table already contains records:
replace_mode = st.checkbox( f":material/sync: Replace Table Mode for `{st.session_state.table_name}`", value=table_exists )
Save to Snowflake
The app creates the database, schema, and table if they don't exist, then inserts each document with metadata including word count and character count.
Full Code
import streamlit as st from pypdf import PdfReader import io import pandas as pd from datetime import datetime try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() st.title(":material/description: Batch Document Text Extractor") st.write("Upload multiple documents at once to extract text and save to Snowflake for RAG applications.") if 'database' not in st.session_state: st.session_state.database = "RAG_DB" if 'schema' not in st.session_state: st.session_state.schema = "RAG_SCHEMA" if 'table_name' not in st.session_state: st.session_state.table_name = "EXTRACTED_DOCUMENTS" with st.container(border=True): st.subheader(":material/analytics: Database Setup") col1, col2, col3 = st.columns(3) with col1: st.session_state.database = st.text_input("Database", value=st.session_state.database, key="db_input") with col2: st.session_state.schema = st.text_input("Schema", value=st.session_state.schema, key="schema_input") with col3: st.session_state.table_name = st.text_input("Table Name", value=st.session_state.table_name, key="table_input") st.info(f":material/location_on: Target location: `{st.session_state.database}.{st.session_state.schema}.{st.session_state.table_name}`") st.caption(":material/lightbulb: Database will be created automatically when you save documents") st.divider() st.subheader(":material/download: Download Review Data") st.write("To get started quickly, download our sample dataset of 100 customer reviews from Avalanche winter sports equipment.") col1, col2 = st.columns([2, 1]) with col1: st.info(":material/info: **Sample Dataset**: 100 customer review files (TXT format) with product feedback, sentiment scores, and order information.") with col2: st.link_button( ":material/download: Download review.zip", "https://github.com/streamlit/30DaysOfAI/raw/refs/heads/main/assets/review.zip", use_container_width=True ) with st.expander(":material/help: How to use the sample data"): st.markdown(""" **Steps:** 1. Click the **Download review.zip** button above 2. Unzip the downloaded file on your computer 3. Use the **Upload Documents** section below to select all 100 review files 4. Click **Extract Text** to process and save to Snowflake **What's included:** - 100 customer review files (`review-001.txt` to `review-100.txt`) - Each review contains: product name, date, review summary, sentiment score, and order ID - Perfect for testing batch processing and building RAG applications **Tip:** You can upload all 100 files at once for optimal batch processing! """) st.divider() st.subheader(":material/upload: Upload Documents") uploaded_files = st.file_uploader( "Choose file(s)", type=["txt", "md", "pdf"], accept_multiple_files=True, help="Supported formats: TXT, MD, PDF. Upload multiple files at once!" ) table_exists = False try: check_result = session.sql(f""" SELECT COUNT(*) as CNT FROM {st.session_state.database}.{st.session_state.schema}.{st.session_state.table_name} """).collect() table_exists = True except: table_exists = False replace_mode = st.checkbox( f":material/sync: Replace Table Mode for `{st.session_state.table_name}`", value=table_exists, help=f"When enabled, clears all existing data in {st.session_state.database}.{st.session_state.schema}.{st.session_state.table_name} before saving new documents" ) if replace_mode: st.warning(f":material/warning: **Replace Mode Enabled** - All existing documents in `{st.session_state.table_name}` will be deleted before saving new ones.") else: st.info(f":material/add: **Append Mode** - New documents will be added to `{st.session_state.table_name}`.") database = st.session_state.database schema = st.session_state.schema table_name = st.session_state.table_name if uploaded_files: with st.container(border=True): st.subheader(":material/upload: Uploaded Documents") st.success(f":material/folder: {len(uploaded_files)} file(s) uploaded") with st.expander(":material/assignment: View Selected Files", expanded=False): file_list_df = pd.DataFrame([ { "File Name": f.name, "Size": f"{f.size:,} bytes", "Type": "TXT" if f.name.lower().endswith('.txt') else "Markdown" if f.name.lower().endswith('.md') else "PDF" if f.name.lower().endswith('.pdf') else "Unknown" } for f in uploaded_files ]) st.dataframe(file_list_df, use_container_width=True) process_button = st.button( f":material/sync: Extract Text from {len(uploaded_files)} File(s)", type="primary", use_container_width=True ) if process_button: success_count = 0 error_count = 0 extracted_data = [] progress_bar = st.progress(0, text="Starting extraction...") status_container = st.empty() for idx, uploaded_file in enumerate(uploaded_files): progress_pct = (idx + 1) / len(uploaded_files) progress_bar.progress(progress_pct, text=f"Processing {idx+1}/{len(uploaded_files)}: {uploaded_file.name}") try: if uploaded_file.name.lower().endswith('.txt'): file_type = "TXT" elif uploaded_file.name.lower().endswith('.md'): file_type = "Markdown" elif uploaded_file.name.lower().endswith('.pdf'): file_type = "PDF" else: file_type = "Unknown" uploaded_file.seek(0) extracted_text = "" if uploaded_file.name.lower().endswith(('.txt', '.md')): extracted_text = uploaded_file.read().decode("utf-8") elif uploaded_file.name.lower().endswith('.pdf'): pdf_reader = PdfReader(io.BytesIO(uploaded_file.read())) for page in pdf_reader.pages: page_text = page.extract_text() if page_text: extracted_text += page_text + "\n\n" if extracted_text and extracted_text.strip(): word_count = len(extracted_text.split()) char_count = len(extracted_text) extracted_data.append({ 'file_name': uploaded_file.name, 'file_type': file_type, 'file_size': uploaded_file.size, 'extracted_text': extracted_text, 'word_count': word_count, 'char_count': char_count }) success_count += 1 else: error_count += 1 status_container.warning(f":material/warning: No text extracted from: {uploaded_file.name}") except Exception as e: error_count += 1 status_container.error(f":material/cancel: Error processing {uploaded_file.name}: {str(e)}") progress_bar.empty() status_container.empty() with st.container(border=True): st.subheader(":material/analytics: Documents Written to a Database Table") col1, col2, col3 = st.columns(3) with col1: st.metric(":material/check_circle: Successful", success_count) with col2: st.metric(":material/cancel: Failed", error_count) with col3: st.metric(":material/analytics: Total Words", f"{sum(d['word_count'] for d in extracted_data):,}") if extracted_data: st.session_state.extracted_data = extracted_data st.success(f":material/check_circle: Successfully extracted text from {success_count} file(s)!") with st.expander(":material/visibility: Preview First 3 Files"): for data in extracted_data[:3]: with st.container(border=True): st.markdown(f"**{data['file_name']}**") st.caption(f"{data['word_count']:,} words") preview_text = data['extracted_text'][:200] if len(data['extracted_text']) > 200: preview_text += "..." st.text(preview_text) if len(extracted_data) > 3: st.caption(f"... and {len(extracted_data) - 3} more") with st.status("Saving to Snowflake...", expanded=True) as status: try: st.write(":material/looks_one: Setting up database structure...") session.sql(f"CREATE DATABASE IF NOT EXISTS {database}").collect() session.sql(f"CREATE SCHEMA IF NOT EXISTS {database}.{schema}").collect() st.write(":material/looks_two: Creating table if needed...") create_table_sql = f""" CREATE TABLE IF NOT EXISTS {database}.{schema}.{table_name} ( DOC_ID NUMBER AUTOINCREMENT, FILE_NAME VARCHAR, FILE_TYPE VARCHAR, FILE_SIZE NUMBER, EXTRACTED_TEXT VARCHAR, UPLOAD_TIMESTAMP TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP(), WORD_COUNT NUMBER, CHAR_COUNT NUMBER ) """ session.sql(create_table_sql).collect() if replace_mode: st.write(":material/sync: Replace mode: Clearing existing data...") try: session.sql(f"TRUNCATE TABLE {database}.{schema}.{table_name}").collect() st.write(" :material/check_circle: Existing data cleared") except Exception as e: st.write(f" :material/warning: No existing data to clear") st.write(f":material/looks_3: Inserting {len(extracted_data)} document(s)...") for idx, data in enumerate(extracted_data, 1): st.caption(f"Saving {idx}/{len(extracted_data)}: {data['file_name']}") safe_text = data['extracted_text'].replace("'", "''") insert_sql = f""" INSERT INTO {database}.{schema}.{table_name} (FILE_NAME, FILE_TYPE, FILE_SIZE, EXTRACTED_TEXT, WORD_COUNT, CHAR_COUNT) VALUES ('{data['file_name']}', '{data['file_type']}', {data['file_size']}, '{safe_text}', {data['word_count']}, {data['char_count']}) """ session.sql(insert_sql).collect() status.update(label=":material/check_circle: All documents saved!", state="complete", expanded=False) mode_msg = "replaced in" if replace_mode else "saved to" st.success(f":material/check_circle: Successfully {mode_msg} `{database}.{schema}.{table_name}`\n\n:material/description: {len(extracted_data)} document(s) now in table") st.session_state.rag_source_table = f"{database}.{schema}.{table_name}" st.session_state.rag_source_database = database st.session_state.rag_source_schema = schema st.balloons() except Exception as e: st.error(f"Error saving to Snowflake: {str(e)}") else: st.warning("No text was successfully extracted from any file.") st.divider() with st.container(border=True): st.subheader(":material/search: View Saved Documents") try: count_result = session.sql(f""" SELECT COUNT(*) as CNT FROM {database}.{schema}.{table_name} """).collect() if count_result: record_count = count_result[0]['CNT'] if record_count > 0: st.warning(f":material/warning: **{record_count} record(s)** currently in table `{database}.{schema}.{table_name}`") else: st.info(":material/inbox: **Table is empty** - No documents uploaded yet.") except: st.info(":material/inbox: **Table doesn't exist yet** - Upload and save documents to create it.") query_button = st.button("Query Table", type="secondary", use_container_width=True) if query_button: try: full_table_name = f"{database}.{schema}.{table_name}" query_sql = f""" SELECT DOC_ID, FILE_NAME, FILE_TYPE, FILE_SIZE, UPLOAD_TIMESTAMP, WORD_COUNT, CHAR_COUNT FROM {full_table_name} ORDER BY UPLOAD_TIMESTAMP DESC """ df = session.sql(query_sql).to_pandas() st.session_state.queried_docs = df st.session_state.full_table_name = full_table_name st.rerun() except Exception as e: st.error(f"Error: {str(e)}") st.info(":material/lightbulb: Table may not exist yet. Upload and save documents first!") if 'queried_docs' in st.session_state and 'full_table_name' in st.session_state: current_full_table_name = f"{st.session_state.database}.{st.session_state.schema}.{st.session_state.table_name}" if st.session_state.full_table_name == current_full_table_name: df = st.session_state.queried_docs if len(df) > 0: st.code(f"{current_full_table_name}", language="sql") col1, col2, col3 = st.columns(3) with col1: st.metric("Documents", len(df)) with col2: st.metric("Words", f"{df['WORD_COUNT'].sum():,}") with col3: st.metric("Characters", f"{df['CHAR_COUNT'].sum():,}") st.divider() st.dataframe( df[['DOC_ID', 'FILE_NAME', 'FILE_TYPE', 'WORD_COUNT', 'UPLOAD_TIMESTAMP']], use_container_width=True ) with st.expander(":material/menu_book: View Full Document Text"): doc_id = st.selectbox( "Select Document ID:", options=df['DOC_ID'].tolist(), format_func=lambda x: f"Doc #{x} - {df[df['DOC_ID']==x]['FILE_NAME'].values[0]}" ) if st.button("Load Text"): text_sql = f"SELECT EXTRACTED_TEXT, FILE_NAME FROM {current_full_table_name} WHERE DOC_ID = {doc_id}" text_result = session.sql(text_sql).to_pandas() if len(text_result) > 0: doc = text_result.iloc[0] st.session_state.loaded_doc_text = doc['EXTRACTED_TEXT'] st.session_state.loaded_doc_name = doc['FILE_NAME'] if 'loaded_doc_text' in st.session_state: st.text_area( st.session_state.loaded_doc_name, value=st.session_state.loaded_doc_text, height=400 ) else: st.info(":material/inbox: Table is empty. Upload files above!") else: st.info(f":material/sync: Showing results for a different table. Click 'Query Table' to refresh.") else: st.info(":material/inbox: No documents queried yet. Click 'Query Table' to view saved documents.") st.divider() st.caption("Day 16: Batch Document Text Extractor for RAG | 30 Days of AI")
Here's the document extraction app in action:
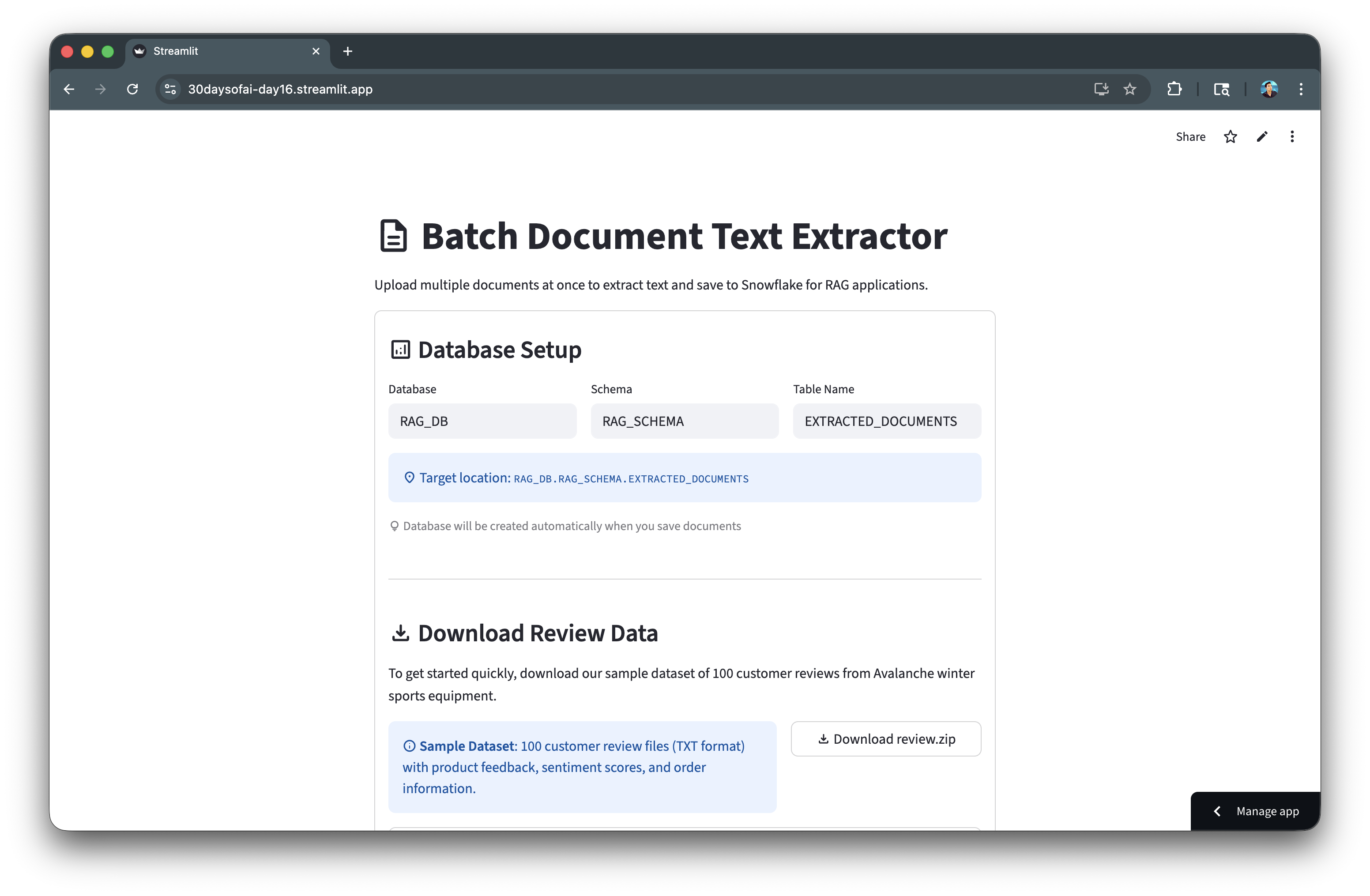
Prepare and Chunk Documents
Text chunking is crucial for effective retrieval. In this section, you'll load the extracted documents from Day 16 and process them into searchable chunks.
Note: This section is repurposed from Day 17 of the #30DaysOfAI learning challenge.
Session State Initialization
The app checks if database configuration was passed from Day 16 via session state. If available, it inherits those values; otherwise, it uses defaults:
if 'day17_database' not in st.session_state: if 'rag_source_database' in st.session_state: st.session_state.day17_database = st.session_state.rag_source_database st.session_state.day17_schema = st.session_state.rag_source_schema st.session_state.day17_table_name = "EXTRACTED_DOCUMENTS" else: st.session_state.day17_database = "RAG_DB" st.session_state.day17_schema = "RAG_SCHEMA" st.session_state.day17_table_name = "EXTRACTED_DOCUMENTS"
Loading Documents
Documents are loaded from Snowflake with a status indicator showing progress. The st.rerun() call ensures the UI updates after loading:
if st.button(":material/folder_open: Load Reviews", type="primary", use_container_width=True): with st.status("Loading reviews from Snowflake...", expanded=True) as status: query = f"""SELECT DOC_ID, FILE_NAME, EXTRACTED_TEXT, WORD_COUNT FROM {st.session_state.day17_database}.{st.session_state.day17_schema}.{st.session_state.day17_table_name}""" df = session.sql(query).to_pandas() st.session_state.loaded_data = df st.rerun()
Processing Strategy
Users choose between keeping each review as a single chunk (recommended for short reviews) or splitting longer reviews with configurable chunk size and overlap:
processing_option = st.radio( "Select processing strategy:", ["Keep each review as a single chunk (Recommended)", "Chunk reviews longer than threshold"] )
Chunking Logic
For chunking, the app uses a sliding window approach with overlap to maintain context between chunks:
for i in range(0, len(words), chunk_size - overlap): chunk_words = words[i:i + chunk_size] chunk_text = ' '.join(chunk_words)
Bulk Insert with write_pandas
The session.write_pandas() method efficiently inserts all chunks in a single operation, with options for replace or append mode.
Full Code
import streamlit as st import pandas as pd import re try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() st.title(":material/sync: Prepare and Chunk Data for RAG") st.write("Load customer reviews from Day 16, process them, and prepare searchable chunks for RAG.") if 'day17_database' not in st.session_state: if 'rag_source_database' in st.session_state: st.session_state.day17_database = st.session_state.rag_source_database st.session_state.day17_schema = st.session_state.rag_source_schema st.session_state.day17_table_name = "EXTRACTED_DOCUMENTS" else: st.session_state.day17_database = "RAG_DB" st.session_state.day17_schema = "RAG_SCHEMA" st.session_state.day17_table_name = "EXTRACTED_DOCUMENTS" if 'day17_chunk_table' not in st.session_state: st.session_state.day17_chunk_table = "REVIEW_CHUNKS" with st.container(border=True): st.subheader(":material/analytics: Source Data Configuration") col1, col2, col3 = st.columns(3) with col1: st.session_state.day17_database = st.text_input( "Database", value=st.session_state.day17_database, key="day17_db_input" ) with col2: st.session_state.day17_schema = st.text_input( "Schema", value=st.session_state.day17_schema, key="day17_schema_input" ) with col3: st.session_state.day17_table_name = st.text_input( "Source Table", value=st.session_state.day17_table_name, key="day17_table_input" ) st.info(f":material/location_on: Loading from: `{st.session_state.day17_database}.{st.session_state.day17_schema}.{st.session_state.day17_table_name}`") st.caption(":material/lightbulb: This should point to the EXTRACTED_DOCUMENTS table from Day 16") if 'loaded_data' in st.session_state: st.success(f":material/check_circle: **{len(st.session_state.loaded_data)} document(s)** already loaded") if st.button(":material/folder_open: Load Reviews", type="primary", use_container_width=True): try: with st.status("Loading reviews from Snowflake...", expanded=True) as status: st.write(":material/wifi: Querying database...") query = f""" SELECT DOC_ID, FILE_NAME, FILE_TYPE, EXTRACTED_TEXT, UPLOAD_TIMESTAMP, WORD_COUNT, CHAR_COUNT FROM {st.session_state.day17_database}.{st.session_state.day17_schema}.{st.session_state.day17_table_name} ORDER BY FILE_NAME """ df = session.sql(query).to_pandas() st.write(f":material/check_circle: Loaded {len(df)} review(s)") status.update(label="Reviews loaded successfully!", state="complete", expanded=False) st.session_state.loaded_data = df st.session_state.source_table = f"{st.session_state.day17_database}.{st.session_state.day17_schema}.{st.session_state.day17_table_name}" st.rerun() except Exception as e: st.error(f"Error loading reviews: {str(e)}") st.info(":material/lightbulb: Make sure you've uploaded review files in Day 16 first!") if 'loaded_data' in st.session_state: with st.container(border=True): st.subheader(":material/looks_one: Review Summary") df = st.session_state.loaded_data col1, col2, col3 = st.columns(3) with col1: st.metric("Total Reviews", len(df)) with col2: st.metric("Total Words", f"{df['WORD_COUNT'].sum():,}") with col3: st.metric("Avg Words/Review", f"{df['WORD_COUNT'].mean():.0f}") st.dataframe(df[['DOC_ID', 'FILE_NAME', 'FILE_TYPE', 'UPLOAD_TIMESTAMP', 'WORD_COUNT']], use_container_width=True) with st.container(border=True): st.subheader(":material/looks_two: Choose Processing Strategy") st.info(""" **Customer Review Processing Options:** Since customer reviews are typically short (~150 words each), you have two options: - **Option 1**: Use each review as-is (Recommended for reviews) - **Option 2**: Chunk longer reviews (For reviews >200 words) """) processing_option = st.radio( "Select processing strategy:", ["Keep each review as a single chunk (Recommended)", "Chunk reviews longer than threshold"], index=0 ) if "Chunk reviews" in processing_option: col1, col2 = st.columns(2) with col1: chunk_size = st.slider( "Chunk Size (words):", min_value=50, max_value=500, value=200, step=50, help="Maximum number of words per chunk" ) with col2: overlap = st.slider( "Overlap (words):", min_value=0, max_value=100, value=50, step=10, help="Number of overlapping words between chunks" ) st.caption(f"Reviews with >{chunk_size} words will be split into chunks of {chunk_size} words with {overlap} word overlap") else: chunk_size = 200 overlap = 50 if st.button(":material/flash_on: Process Reviews", type="primary", use_container_width=True): chunks = [] with st.status("Processing reviews...", expanded=True) as status: if "Keep each review" in processing_option: st.write(":material/edit_note: Creating one chunk per review...") for idx, row in df.iterrows(): chunks.append({ 'doc_id': row['DOC_ID'], 'file_name': row['FILE_NAME'], 'chunk_id': idx + 1, 'chunk_text': row['EXTRACTED_TEXT'], 'chunk_size': row['WORD_COUNT'], 'chunk_type': 'full_review' }) st.write(f":material/check_circle: Created {len(chunks)} chunks (1 per review)") else: st.write(f":material/edit_note: Chunking reviews longer than {chunk_size} words...") chunk_id = 1 for idx, row in df.iterrows(): text = row['EXTRACTED_TEXT'] words = text.split() if len(words) <= chunk_size: chunks.append({ 'doc_id': row['DOC_ID'], 'file_name': row['FILE_NAME'], 'chunk_id': chunk_id, 'chunk_text': text, 'chunk_size': len(words), 'chunk_type': 'full_review' }) chunk_id += 1 else: for i in range(0, len(words), chunk_size - overlap): chunk_words = words[i:i + chunk_size] chunk_text = ' '.join(chunk_words) chunks.append({ 'doc_id': row['DOC_ID'], 'file_name': row['FILE_NAME'], 'chunk_id': chunk_id, 'chunk_text': chunk_text, 'chunk_size': len(chunk_words), 'chunk_type': 'chunked_review' }) chunk_id += 1 st.write(f":material/check_circle: Created {len(chunks)} chunks from {len(df)} reviews") status.update(label="Processing complete!", state="complete", expanded=False) st.session_state.review_chunks = chunks st.session_state.processing_option = processing_option st.success(f":material/check_circle: Processed {len(df)} reviews into {len(chunks)} searchable chunks!") if 'review_chunks' in st.session_state: with st.container(border=True): st.subheader(":material/looks_3: Processed Review Chunks") chunks = st.session_state.review_chunks col1, col2, col3 = st.columns(3) with col1: st.metric("Total Chunks", len(chunks)) with col2: full_reviews = len([c for c in chunks if c['chunk_type'] == 'full_review']) st.metric("Full Reviews", full_reviews) with col3: split_reviews = len([c for c in chunks if c['chunk_type'] == 'chunked_review']) st.metric("Split Reviews", split_reviews) with st.expander(":material/description: View Chunks"): chunks_df = pd.DataFrame(chunks) st.dataframe(chunks_df[['chunk_id', 'file_name', 'chunk_size', 'chunk_type', 'chunk_text']], use_container_width=True) with st.container(border=True): st.subheader(":material/looks_4: Save Chunks to Snowflake") chunks = st.session_state.review_chunks col1, col2 = st.columns([2, 1]) with col1: st.session_state.day17_chunk_table = st.text_input( "Chunk Table Name", value=st.session_state.day17_chunk_table, help="Table name for storing review chunks", key="day17_chunk_table_input" ) full_chunk_table = f"{st.session_state.day17_database}.{st.session_state.day17_schema}.{st.session_state.day17_chunk_table}" st.code(full_chunk_table, language="sql") chunk_table_exists = False try: count_result = session.sql(f""" SELECT COUNT(*) as CNT FROM {full_chunk_table} """).collect() if count_result: record_count = count_result[0]['CNT'] if record_count > 0: st.warning(f":material/warning: **{record_count} chunk(s)** currently in table `{full_chunk_table}`") chunk_table_exists = True else: st.info(":material/inbox: **Chunk table is empty** - No chunks saved yet.") chunk_table_exists = False except: st.info(":material/inbox: **Chunk table doesn't exist yet** - Will be created when you save chunks.") chunk_table_exists = False if 'day17_replace_mode' not in st.session_state: st.session_state.day17_replace_mode = chunk_table_exists else: if 'day17_last_chunk_table' not in st.session_state or st.session_state.day17_last_chunk_table != full_chunk_table: st.session_state.day17_replace_mode = chunk_table_exists st.session_state.day17_last_chunk_table = full_chunk_table replace_mode = st.checkbox( f":material/sync: Replace Table Mode for `{st.session_state.day17_chunk_table}`", help=f"When enabled, clears all existing data in {full_chunk_table} before saving new chunks", key="day17_replace_mode" ) if replace_mode: st.warning("**Replace Mode Active**: Existing chunks will be deleted before saving new ones.") else: st.success("**Append Mode Active**: New chunks will be added to existing data.") if st.button(":material/save: Save Chunks to Snowflake", type="primary", use_container_width=True): try: with st.status("Saving chunks to Snowflake...", expanded=True) as status: st.write(":material/looks_one: Checking table...") create_table_sql = f""" CREATE TABLE IF NOT EXISTS {full_chunk_table} ( CHUNK_ID NUMBER, DOC_ID NUMBER, FILE_NAME VARCHAR, CHUNK_TEXT VARCHAR, CHUNK_SIZE NUMBER, CHUNK_TYPE VARCHAR, CREATED_TIMESTAMP TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP() ) """ session.sql(create_table_sql).collect() if replace_mode: st.write(":material/sync: Replace mode: Clearing existing chunks...") try: session.sql(f"TRUNCATE TABLE {full_chunk_table}").collect() st.write(" :material/check_circle: Existing chunks cleared") except Exception as e: st.write(f" :material/warning: No existing chunks to clear") st.write(f":material/looks_3: Inserting {len(chunks)} chunk(s)...") chunks_df = pd.DataFrame(chunks) chunks_df_upper = chunks_df[['chunk_id', 'doc_id', 'file_name', 'chunk_text', 'chunk_size', 'chunk_type']].copy() chunks_df_upper.columns = ['CHUNK_ID', 'DOC_ID', 'FILE_NAME', 'CHUNK_TEXT', 'CHUNK_SIZE', 'CHUNK_TYPE'] if replace_mode: session.write_pandas(chunks_df_upper, table_name=st.session_state.day17_chunk_table, database=st.session_state.day17_database, schema=st.session_state.day17_schema, overwrite=True) else: session.write_pandas(chunks_df_upper, table_name=st.session_state.day17_chunk_table, database=st.session_state.day17_database, schema=st.session_state.day17_schema, overwrite=False) status.update(label=":material/check_circle: Chunks saved!", state="complete", expanded=False) mode_msg = "replaced in" if replace_mode else "saved to" st.success(f":material/check_circle: Successfully {mode_msg} `{full_chunk_table}`\n\n:material/description: {len(chunks)} chunk(s) now in table") st.session_state.chunks_table = full_chunk_table st.session_state.chunks_database = st.session_state.day17_database st.session_state.chunks_schema = st.session_state.day17_schema st.session_state.chunk_table_saved = True st.balloons() except Exception as e: st.error(f"Error saving chunks: {str(e)}") with st.container(border=True): st.subheader(":material/search: View Saved Chunks") full_chunk_table = f"{st.session_state.day17_database}.{st.session_state.day17_schema}.{st.session_state.day17_chunk_table}" st.caption(f":material/analytics: Querying chunk table: `{full_chunk_table}`") query_button = st.button(":material/analytics: Query Chunk Table", type="secondary", use_container_width=True) if query_button: try: query_sql = f""" SELECT CHUNK_ID, FILE_NAME, CHUNK_SIZE, CHUNK_TYPE, LEFT(CHUNK_TEXT, 100) AS TEXT_PREVIEW, CREATED_TIMESTAMP FROM {full_chunk_table} ORDER BY CHUNK_ID """ chunks_df = session.sql(query_sql).to_pandas() st.session_state.queried_chunks = chunks_df st.session_state.queried_chunks_table = full_chunk_table st.rerun() except Exception as e: st.error(f"Error querying chunks: {str(e)}") if 'queried_chunks' in st.session_state and st.session_state.get('queried_chunks_table') == full_chunk_table: chunks_df = st.session_state.queried_chunks if len(chunks_df) > 0: st.code(full_chunk_table, language="sql") col1, col2, col3 = st.columns(3) with col1: st.metric("Total Chunks", len(chunks_df)) with col2: full_count = len(chunks_df[chunks_df['CHUNK_TYPE'] == 'full_review']) st.metric("Full Reviews", full_count) with col3: split_count = len(chunks_df[chunks_df['CHUNK_TYPE'] == 'chunked_review']) st.metric("Split Reviews", split_count) st.dataframe( chunks_df[['CHUNK_ID', 'FILE_NAME', 'CHUNK_SIZE', 'CHUNK_TYPE', 'TEXT_PREVIEW']], use_container_width=True ) with st.expander(":material/menu_book: View Full Chunk Text"): chunk_id = st.selectbox( "Select Chunk ID:", options=chunks_df['CHUNK_ID'].tolist(), format_func=lambda x: f"Chunk #{x} - {chunks_df[chunks_df['CHUNK_ID']==x]['FILE_NAME'].values[0]}", key="chunk_text_selector" ) if st.button("Load Chunk Text", key="load_chunk_text_btn"): st.session_state.selected_chunk_id = chunk_id st.session_state.load_chunk_text = True st.rerun() if st.session_state.get('load_chunk_text') and st.session_state.get('selected_chunk_id'): text_sql = f"SELECT CHUNK_TEXT, FILE_NAME FROM {full_chunk_table} WHERE CHUNK_ID = {st.session_state.selected_chunk_id}" text_result = session.sql(text_sql).to_pandas() if len(text_result) > 0: chunk = text_result.iloc[0] st.text_area( chunk['FILE_NAME'], value=chunk['CHUNK_TEXT'], height=300, key=f"chunk_text_display_{st.session_state.selected_chunk_id}" ) else: st.info(":material/inbox: No chunks found in table.") else: st.info(":material/inbox: No chunks queried yet. Click 'Query Chunk Table' to view saved chunks.") st.divider() st.caption("Day 17: Loading and Transforming Customer Reviews for RAG | 30 Days of AI")
Here's the chunking process in action:
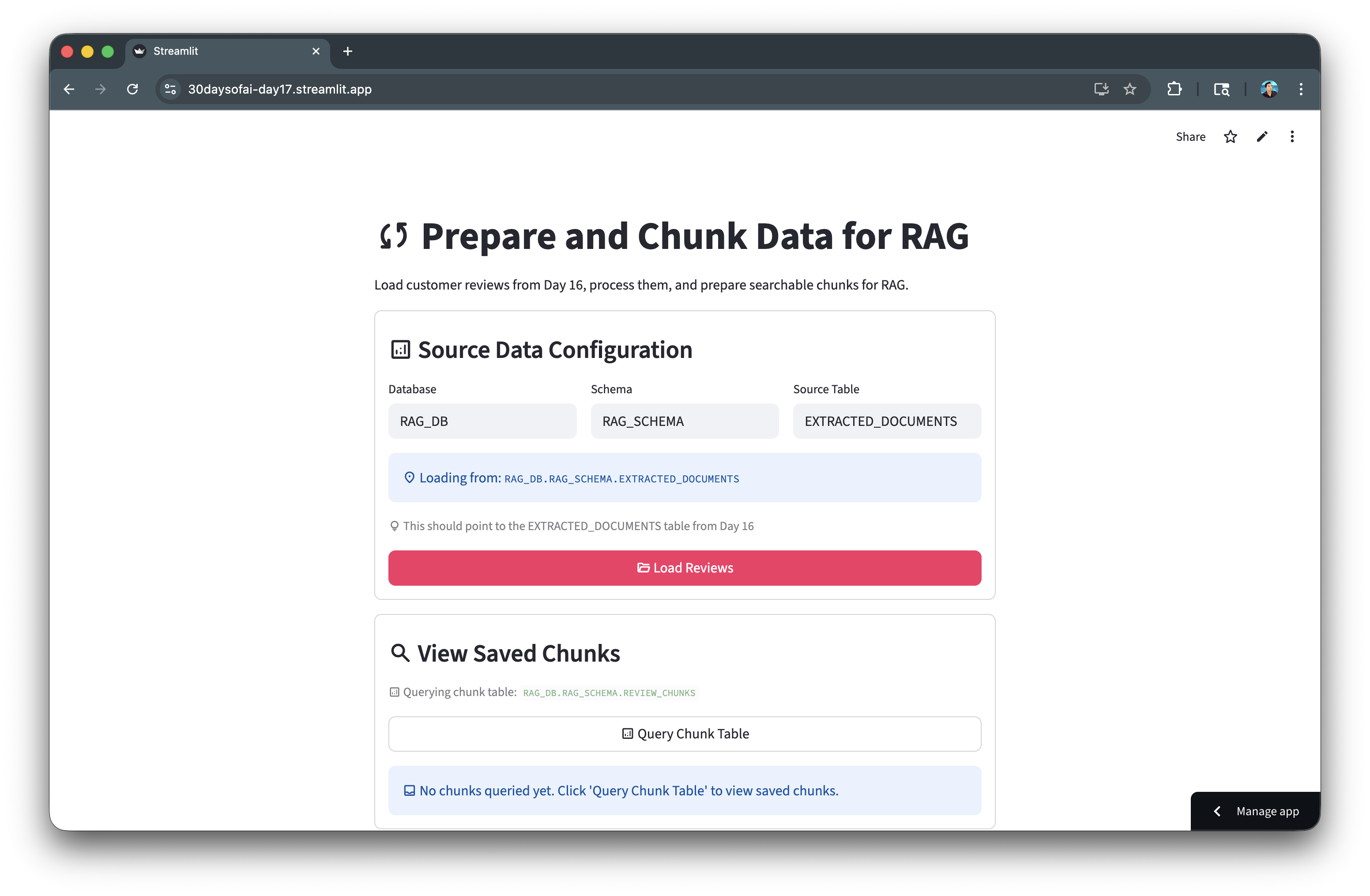
Generate Embeddings
Embeddings convert text into numerical vectors that capture semantic meaning, enabling similarity-based search. In this section, you'll generate 768-dimensional embeddings for your chunked reviews using Snowflake Cortex.
Note: This section is repurposed from Day 18 of the #30DaysOfAI learning challenge.
Understanding Embeddings
Embeddings transform text into dense numerical vectors where similar meanings produce similar vectors. The embed_text_768 function from Snowflake Cortex generates 768-dimensional vectors using the snowflake-arctic-embed-m model.
from snowflake.cortex import embed_text_768 emb = embed_text_768(model='snowflake-arctic-embed-m', text=row['CHUNK_TEXT'])
Storing Embeddings with VECTOR Type
Snowflake's native VECTOR(FLOAT, 768) data type efficiently stores embedding vectors:
create_table_sql = f""" CREATE OR REPLACE TABLE {full_embedding_table} ( CHUNK_ID NUMBER, EMBEDDING VECTOR(FLOAT, 768), CREATED_TIMESTAMP TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP() ) """ session.sql(create_table_sql).collect()
Batch Processing with Progress Tracking
Processing embeddings in batches prevents timeouts and provides user feedback:
for i in range(0, total_chunks, batch_size): batch_end = min(i + batch_size, total_chunks) st.write(f"Processing chunks {i+1} to {batch_end} of {total_chunks}...") for idx, row in df.iloc[i:batch_end].iterrows(): emb = embed_text_768(model='snowflake-arctic-embed-m', text=row['CHUNK_TEXT']) embeddings.append({'chunk_id': row['CHUNK_ID'], 'embedding': emb}) progress_bar.progress(batch_end / total_chunks)
Inserting Vector Data
Convert embeddings to Snowflake's vector format for storage:
emb_array = "[" + ",".join([str(float(x)) for x in emb_list]) + "]" insert_sql = f""" INSERT INTO {full_embedding_table} (CHUNK_ID, EMBEDDING) SELECT {emb_data['chunk_id']}, {emb_array}::VECTOR(FLOAT, 768) """ session.sql(insert_sql).collect()
Full Code
Copy and paste this code into day18.py:
# Day 18 # Generating Embeddings for Customer Reviews import streamlit as st from snowflake.cortex import embed_text_768 import pandas as pd import numpy as np st.title(":material/calculate: Embeddings Generator for Customer Reviews") st.write("Generate embeddings for review chunks from Day 17 to enable semantic search.") # Connect to Snowflake try: # Works in Streamlit in Snowflake from snowflake.snowpark.context import get_active_session session = get_active_session() except: # Works locally and on Streamlit Community Cloud from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() # Initialize session state for database configuration if 'day18_database' not in st.session_state: # Check if we have chunks from Day 17 if 'chunks_database' in st.session_state: st.session_state.day18_database = st.session_state.chunks_database st.session_state.day18_schema = st.session_state.chunks_schema st.session_state.day18_chunk_table = "REVIEW_CHUNKS" else: st.session_state.day18_database = "RAG_DB" st.session_state.day18_schema = "RAG_SCHEMA" st.session_state.day18_chunk_table = "REVIEW_CHUNKS" if 'day18_embedding_table' not in st.session_state: st.session_state.day18_embedding_table = "REVIEW_EMBEDDINGS" # Explanation with st.expander(":material/library_books: What are embeddings?", expanded=True): st.markdown(""" **Embeddings** convert text into numbers (vectors) that capture meaning: - Similar texts → Similar vectors - Different texts → Different vectors - Enables "search by meaning" (semantic search) The model outputs **768 numbers** for any text input. **In RAG for Customer Reviews**: Each review (or chunk) gets its own embedding, allowing semantic search to find relevant customer feedback! **Example**: Search for "warm gloves" will find reviews mentioning "provides good warmth", "kept hands toasty", even without exact keywords! """) # Source Data Configuration and Load Section with st.container(border=True): st.subheader(":material/analytics: Source Data Configuration") # Database configuration col1, col2, col3 = st.columns(3) with col1: st.session_state.day18_database = st.text_input( "Database", value=st.session_state.day18_database, key="day18_db_input" ) with col2: st.session_state.day18_schema = st.text_input( "Schema", value=st.session_state.day18_schema, key="day18_schema_input" ) with col3: st.session_state.day18_chunk_table = st.text_input( "Chunks Table", value=st.session_state.day18_chunk_table, key="day18_chunk_table_input" ) st.info(f":material/location_on: Loading from: `{st.session_state.day18_database}.{st.session_state.day18_schema}.{st.session_state.day18_chunk_table}`") st.caption(":material/lightbulb: This should point to the REVIEW_CHUNKS table from Day 17") # Check for existing loaded data if 'chunks_data' in st.session_state: st.success(f":material/check_circle: **{len(st.session_state.chunks_data)} chunk(s)** already loaded") # Load chunks button if st.button(":material/folder_open: Load Chunks", type="primary", use_container_width=True): try: with st.status("Loading chunks...", expanded=True) as status: st.write(":material/wifi: Querying database...") query = f""" SELECT CHUNK_ID, DOC_ID, FILE_NAME, CHUNK_TEXT, CHUNK_SIZE, CHUNK_TYPE FROM {st.session_state.day18_database}.{st.session_state.day18_schema}.{st.session_state.day18_chunk_table} ORDER BY CHUNK_ID """ df = session.sql(query).to_pandas() st.write(f":material/check_circle: Loaded {len(df)} chunks") status.update(label="Chunks loaded successfully!", state="complete", expanded=False) # Store in session state st.session_state.chunks_data = df st.rerun() except Exception as e: st.error(f"Error loading chunks: {str(e)}") st.info(":material/lightbulb: Make sure you've processed reviews in Day 17 first!") # Main content - Chunk Summary if 'chunks_data' in st.session_state: with st.container(border=True): st.subheader(":material/looks_one: Chunk Summary") df = st.session_state.chunks_data col1, col2, col3 = st.columns(3) with col1: st.metric("Total Chunks", len(df)) with col2: st.metric("Unique Reviews", df['FILE_NAME'].nunique()) with col3: st.metric("Avg Chunk Size", f"{df['CHUNK_SIZE'].mean():.0f} words") # Show chunk type distribution if 'CHUNK_TYPE' in df.columns: st.write("**Chunk Type Distribution:**") chunk_type_counts = df['CHUNK_TYPE'].value_counts() col1, col2 = st.columns(2) with col1: st.metric("Full Reviews", chunk_type_counts.get('full_review', 0)) with col2: st.metric("Split Reviews", chunk_type_counts.get('chunked_review', 0)) with st.expander(":material/description: Preview Chunks"): st.dataframe(df.head(10), use_container_width=True) # Generate embeddings with st.container(border=True): st.subheader(":material/looks_two: Generate Embeddings") st.info(""" **What happens here:** - Each review chunk is converted to a 768-dimensional vector - Embeddings are stored in Snowflake for semantic search - Enables finding relevant reviews based on meaning, not just keywords **For Customer Reviews**: This allows your RAG system to: - Find reviews about "durability" even if they mention "long-lasting" or "fell apart" - Search for "warm" products and find "toasty", "cold hands", "insulation" - Group similar feedback together semantically """) # Batch size selection batch_size = st.selectbox("Batch Size", [10, 25, 50, 100], index=1, help="Number of chunks to process at once") if st.button(":material/calculate: Generate Embeddings", type="primary", use_container_width=True): try: with st.status("Generating embeddings...", expanded=True) as status: embeddings = [] total_chunks = len(df) progress_bar = st.progress(0) for i in range(0, total_chunks, batch_size): batch_end = min(i + batch_size, total_chunks) st.write(f"Processing chunks {i+1} to {batch_end} of {total_chunks}...") for idx, row in df.iloc[i:batch_end].iterrows(): # Generate embedding using the correct function signature emb = embed_text_768(model='snowflake-arctic-embed-m', text=row['CHUNK_TEXT']) embeddings.append({ 'chunk_id': row['CHUNK_ID'], 'embedding': emb }) # Update progress progress = batch_end / total_chunks progress_bar.progress(progress) status.update(label="Embeddings generated!", state="complete", expanded=False) # Store in session state st.session_state.embeddings_data = embeddings st.success(f":material/check_circle: Generated {len(embeddings)} embeddings for {len(df)} review chunks!") except Exception as e: st.error(f"Error generating embeddings: {str(e)}") # View embeddings if 'embeddings_data' in st.session_state: with st.container(border=True): st.subheader(":material/looks_3: View Embeddings") embeddings = st.session_state.embeddings_data col1, col2 = st.columns(2) with col1: st.metric("Embeddings Generated", len(embeddings)) with col2: st.metric("Dimensions per Embedding", 768) # Show sample embedding with st.expander(":material/search: View Sample Embedding"): sample_emb = embeddings[0]['embedding'] st.write("**First 10 values:**") st.write(sample_emb[:10]) # Save embeddings to Snowflake with st.container(border=True): st.subheader(":material/looks_4: Save Embeddings to Snowflake") embeddings = st.session_state.embeddings_data # Embedding table name col1, col2 = st.columns([2, 1]) with col1: st.session_state.day18_embedding_table = st.text_input( "Embeddings Table Name", value=st.session_state.day18_embedding_table, help="Table name for storing embeddings", key="day18_embedding_table_input" ) full_embedding_table = f"{st.session_state.day18_database}.{st.session_state.day18_schema}.{st.session_state.day18_embedding_table}" st.code(full_embedding_table, language="sql") # Check if embeddings table exists and show status try: check_query = f""" SELECT COUNT(*) as count FROM {full_embedding_table} """ result = session.sql(check_query).collect() current_count = result[0]['COUNT'] if current_count > 0: st.warning(f":material/warning: **{current_count:,} embedding(s)** currently in table `{full_embedding_table}`") embedding_table_exists = True else: st.info(":material/inbox: **Embedding table is empty** - No embeddings saved yet.") embedding_table_exists = False except: st.info(":material/inbox: **Embedding table doesn't exist yet** - Will be created when you save embeddings.") embedding_table_exists = False # Initialize or update checkbox state based on table status if 'day18_replace_mode' not in st.session_state: st.session_state.day18_replace_mode = embedding_table_exists else: if 'day18_last_embedding_table' not in st.session_state or st.session_state.day18_last_embedding_table != full_embedding_table: st.session_state.day18_replace_mode = embedding_table_exists st.session_state.day18_last_embedding_table = full_embedding_table # Replace mode checkbox replace_mode = st.checkbox( f":material/sync: Replace Table Mode for `{st.session_state.day18_embedding_table}`", help=f"When enabled, replaces all existing embeddings in {full_embedding_table}", key="day18_replace_mode" ) if replace_mode: st.warning("**Replace Mode Active**: Existing embeddings will be deleted before saving new ones.") else: st.success("**Append Mode Active**: New embeddings will be added to existing data.") if st.button(":material/save: Save Embeddings to Snowflake", type="primary", use_container_width=True): try: with st.status("Saving embeddings...", expanded=True) as status: # Step 1: Create or truncate embeddings table st.write(":material/looks_one: Preparing table...") if replace_mode: # Replace existing data create_table_sql = f""" CREATE OR REPLACE TABLE {full_embedding_table} ( CHUNK_ID NUMBER, EMBEDDING VECTOR(FLOAT, 768), CREATED_TIMESTAMP TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP() ) """ session.sql(create_table_sql).collect() st.write(":material/check_circle: Replaced existing table") else: # Create if doesn't exist create_table_sql = f""" CREATE TABLE IF NOT EXISTS {full_embedding_table} ( CHUNK_ID NUMBER, EMBEDDING VECTOR(FLOAT, 768), CREATED_TIMESTAMP TIMESTAMP_NTZ DEFAULT CURRENT_TIMESTAMP() ) """ session.sql(create_table_sql).collect() st.write(":material/check_circle: Table ready") # Step 2: Insert embeddings st.write(f":material/looks_two: Inserting {len(embeddings)} embeddings...") for i, emb_data in enumerate(embeddings): # Get embedding list if isinstance(emb_data['embedding'], list): emb_list = emb_data['embedding'] else: emb_list = list(emb_data['embedding']) # Convert to proper array format for Snowflake emb_array = "[" + ",".join([str(float(x)) for x in emb_list]) + "]" insert_sql = f""" INSERT INTO {full_embedding_table} (CHUNK_ID, EMBEDDING) SELECT {emb_data['chunk_id']}, {emb_array}::VECTOR(FLOAT, 768) """ session.sql(insert_sql).collect() if (i + 1) % 10 == 0: st.write(f"Saved {i + 1} of {len(embeddings)} embeddings...") status.update(label="Embeddings saved!", state="complete", expanded=False) mode_msg = "replaced in" if replace_mode else "saved to" st.success(f":material/check_circle: Successfully {mode_msg} `{full_embedding_table}`\n\n:material/calculate: {len(embeddings)} embedding(s) now in table") # Store for Day 19 st.session_state.embeddings_table = full_embedding_table st.session_state.embeddings_database = st.session_state.day18_database st.session_state.embeddings_schema = st.session_state.day18_schema st.balloons() except Exception as e: st.error(f"Error saving embeddings: {str(e)}") # View Saved Embeddings Section with st.container(border=True): st.subheader(":material/search: View Saved Embeddings") # Check if embeddings table exists and show record count full_embedding_table = f"{st.session_state.day18_database}.{st.session_state.day18_schema}.{st.session_state.day18_embedding_table}" try: count_result = session.sql(f""" SELECT COUNT(*) as CNT FROM {full_embedding_table} """).collect() if count_result: record_count = count_result[0]['CNT'] if record_count > 0: st.warning(f":material/warning: **{record_count:,} embedding(s)** currently in table `{full_embedding_table}`") else: st.info(":material/inbox: **Embedding table is empty** - Generate and save embeddings above.") except: st.info(":material/inbox: **Embedding table doesn't exist yet** - Generate and save embeddings to create it.") query_button = st.button(":material/analytics: Query Embedding Table", type="secondary", use_container_width=True) if query_button: try: query = f""" SELECT CHUNK_ID, EMBEDDING, CREATED_TIMESTAMP, VECTOR_L2_DISTANCE(EMBEDDING, EMBEDDING) as SELF_DISTANCE FROM {full_embedding_table} ORDER BY CHUNK_ID """ result_df = session.sql(query).to_pandas() # Store in session state st.session_state.queried_embeddings = result_df st.session_state.queried_embeddings_table = full_embedding_table st.rerun() except Exception as e: st.error(f"Error querying embeddings: {str(e)}") # Display results if available in session state if 'queried_embeddings' in st.session_state and st.session_state.get('queried_embeddings_table') == full_embedding_table: emb_df = st.session_state.queried_embeddings if len(emb_df) > 0: st.code(full_embedding_table, language="sql") # Summary metrics col1, col2 = st.columns(2) with col1: st.metric("Total Embeddings", len(emb_df)) with col2: st.metric("Dimensions", "768") # Display table without the EMBEDDING column for readability # Check which columns exist (case-insensitive) embedding_col = None for col in emb_df.columns: if col.upper() == 'EMBEDDING': embedding_col = col break if embedding_col: display_df = emb_df.drop(columns=[embedding_col]) else: display_df = emb_df st.dataframe(display_df, use_container_width=True) st.info(":material/lightbulb: Self-distance should be 0, confirming embeddings are stored correctly") # View individual embedding vectors (only if EMBEDDING column exists) if embedding_col: with st.expander(":material/search: View Individual Embedding Vectors"): st.write("Select a CHUNK_ID to view its full 768-dimensional embedding vector:") # Find CHUNK_ID column (case-insensitive) chunk_id_col = None for col in emb_df.columns: if col.upper() == 'CHUNK_ID': chunk_id_col = col break chunk_ids = emb_df[chunk_id_col].tolist() selected_chunk = st.selectbox("Select CHUNK_ID", chunk_ids, key="view_embedding_chunk") if st.button(":material/analytics: Load Embedding Vector", key="load_embedding_btn"): # Get the embedding for selected chunk selected_emb = emb_df[emb_df[chunk_id_col] == selected_chunk][embedding_col].iloc[0] # Store in session state st.session_state.loaded_embedding = selected_emb st.session_state.loaded_embedding_chunk = selected_chunk st.rerun() # Display loaded embedding if 'loaded_embedding' in st.session_state: st.write(f"**Embedding Vector for CHUNK_ID {st.session_state.loaded_embedding_chunk}:**") # Convert to list if needed emb_vector = st.session_state.loaded_embedding if isinstance(emb_vector, str): # If it's a string representation, parse it import json emb_vector = json.loads(emb_vector) elif hasattr(emb_vector, 'tolist'): emb_vector = emb_vector.tolist() elif not isinstance(emb_vector, list): emb_vector = list(emb_vector) st.caption(f"Vector length: {len(emb_vector)} dimensions") # Display the full embedding vector as code st.code(emb_vector, language="python") else: st.info(":material/inbox: No embeddings found in table.") else: st.info(":material/inbox: No embeddings queried yet. Click 'Query Embedding Table' to view saved embeddings.") st.divider() st.caption("Day 18: Generating Embeddings for Customer Reviews | 30 Days of AI")
Here's the embedding generator in action:
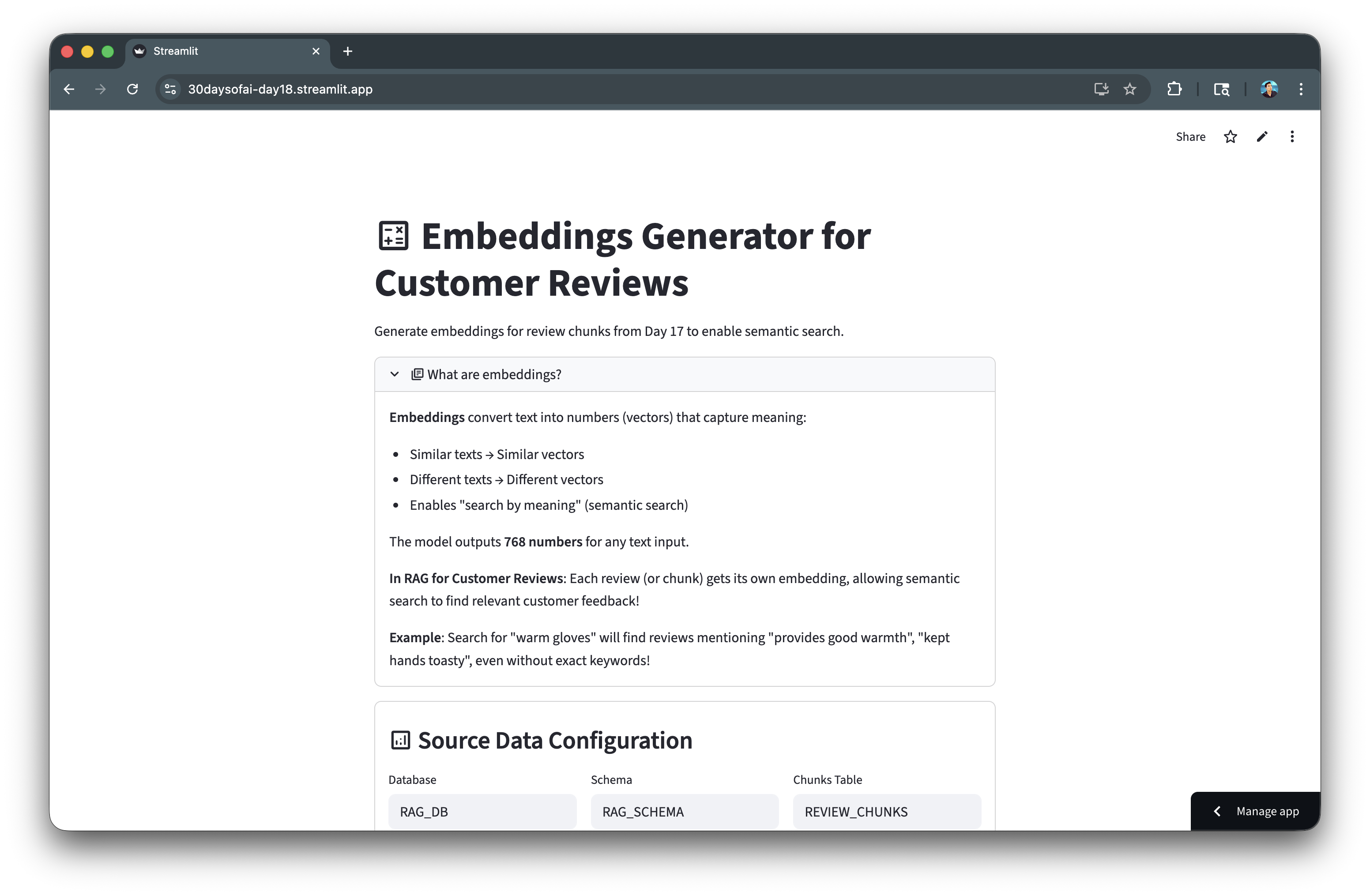
Create Cortex Search Service
Cortex Search provides semantic search over your chunked data. In this section, you'll create a search service that indexes your chunks for fast retrieval.
Note: This section is repurposed from Day 19 of the #30DaysOfAI learning challenge.
Service Configuration
The CREATE CORTEX SEARCH SERVICE statement defines how your data is indexed:
- ON: Specifies the text column to search (
CHUNK_TEXT) - ATTRIBUTES: Additional columns returned with results (
FILE_NAME,CHUNK_TYPE,CHUNK_ID) - TARGET_LAG: How often the index refreshes (1 hour keeps it reasonably fresh)
- WAREHOUSE: The compute resource used for indexing
Key Configuration Options
| Parameter | Description |
|---|---|
ON | The text column to search |
ATTRIBUTES | Metadata columns to return |
TARGET_LAG | How often to refresh the index |
WAREHOUSE | Compute resource for indexing |
Full Code
CREATE OR REPLACE CORTEX SEARCH SERVICE RAG_DB.RAG_SCHEMA.CUSTOMER_REVIEW_SEARCH ON CHUNK_TEXT ATTRIBUTES FILE_NAME, CHUNK_TYPE, CHUNK_ID WAREHOUSE = COMPUTE_WH TARGET_LAG = '1 hour' AS ( SELECT CHUNK_ID, FILE_NAME, CHUNK_TYPE, CHUNK_TEXT FROM RAG_DB.RAG_SCHEMA.REVIEW_CHUNKS );
Verify your service was created:
SHOW CORTEX SEARCH SERVICES IN SCHEMA RAG_DB.RAG_SCHEMA;
Here's the Cortex Search service creation:
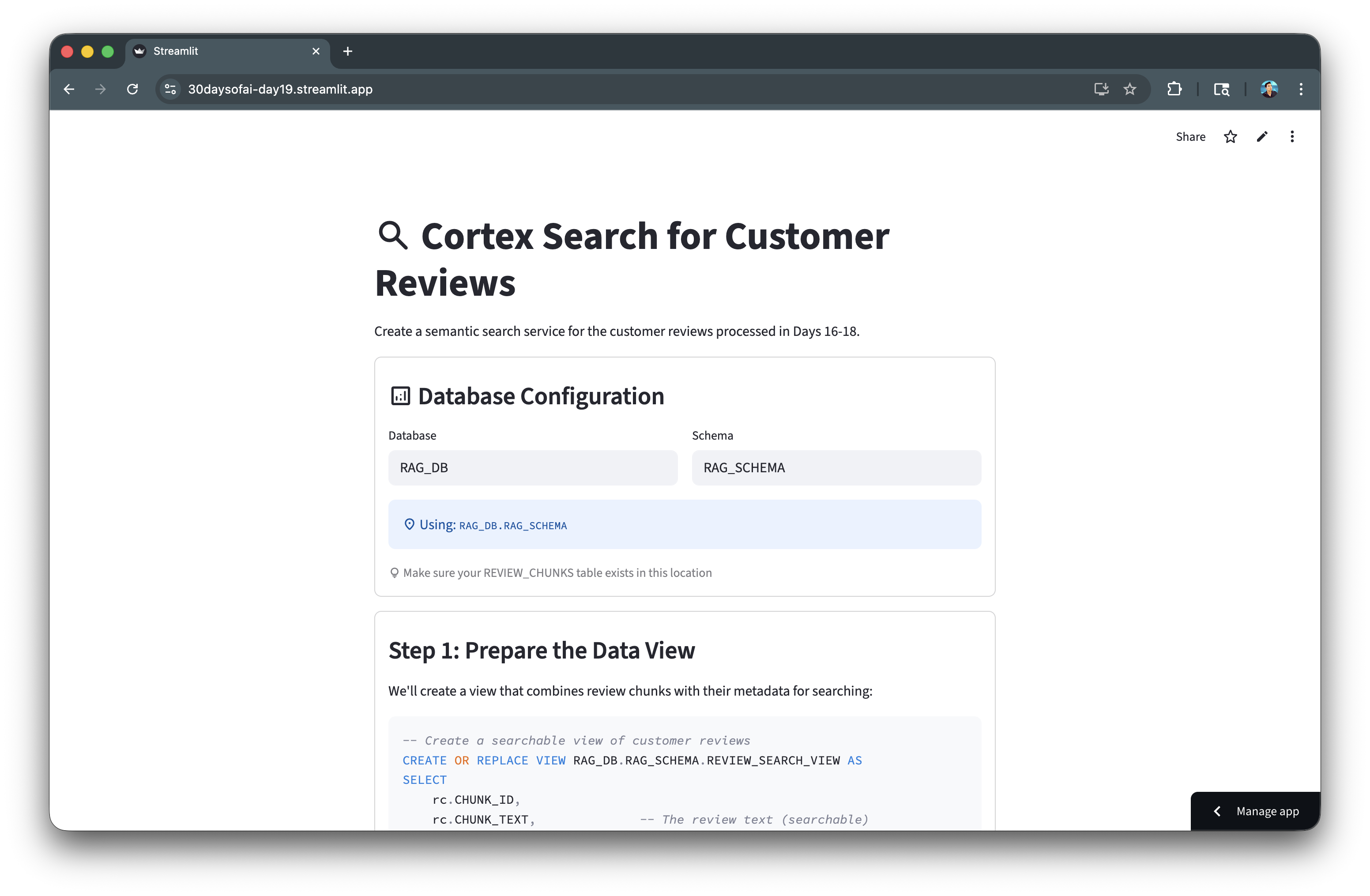
Query Cortex Search
Use the Python SDK to search your documents semantically. This section builds an interactive search interface.
Note: This section is repurposed from Day 20 of the #30DaysOfAI learning challenge.
Cortex Search Python SDK
The snowflake.core.Root API provides programmatic access to Cortex Search. Navigate the object hierarchy using database, schema, and service name:
from snowflake.core import Root root = Root(session) parts = service_path.split(".") svc = root.databases[parts[0]].schemas[parts[1]].cortex_search_services[parts[2]]
Search Method
The search() method accepts a natural language query and returns relevant chunks ranked by semantic similarity:
results = svc.search( query="warm thermal gloves", columns=["CHUNK_TEXT", "FILE_NAME", "CHUNK_TYPE", "CHUNK_ID"], limit=5 )
Displaying Results
Each result is displayed in a bordered container with file metadata and chunk text for easy scanning.
Full Code
Copy and paste this code into day20.py:
import streamlit as st from snowflake.core import Root try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() st.title(":material/search: Querying Cortex Search") search_service = st.text_input( "Search Service:", value="RAG_DB.RAG_SCHEMA.CUSTOMER_REVIEW_SEARCH" ) query = st.text_input("Enter your search query:", value="warm thermal gloves") num_results = st.slider("Number of results:", 1, 20, 5) if st.button(":material/search: Search", type="primary"): if query and search_service: root = Root(session) parts = search_service.split(".") svc = root.databases[parts[0]].schemas[parts[1]].cortex_search_services[parts[2]] with st.spinner("Searching..."): results = svc.search( query=query, columns=["CHUNK_TEXT", "FILE_NAME", "CHUNK_TYPE", "CHUNK_ID"], limit=num_results ) st.success(f":material/check_circle: Found {len(results.results)} result(s)!") for i, item in enumerate(results.results, 1): with st.container(border=True): st.markdown(f"**Result {i}** - {item.get('FILE_NAME', 'N/A')}") st.write(item.get("CHUNK_TEXT", "No text found"))
Here's the search interface in action:
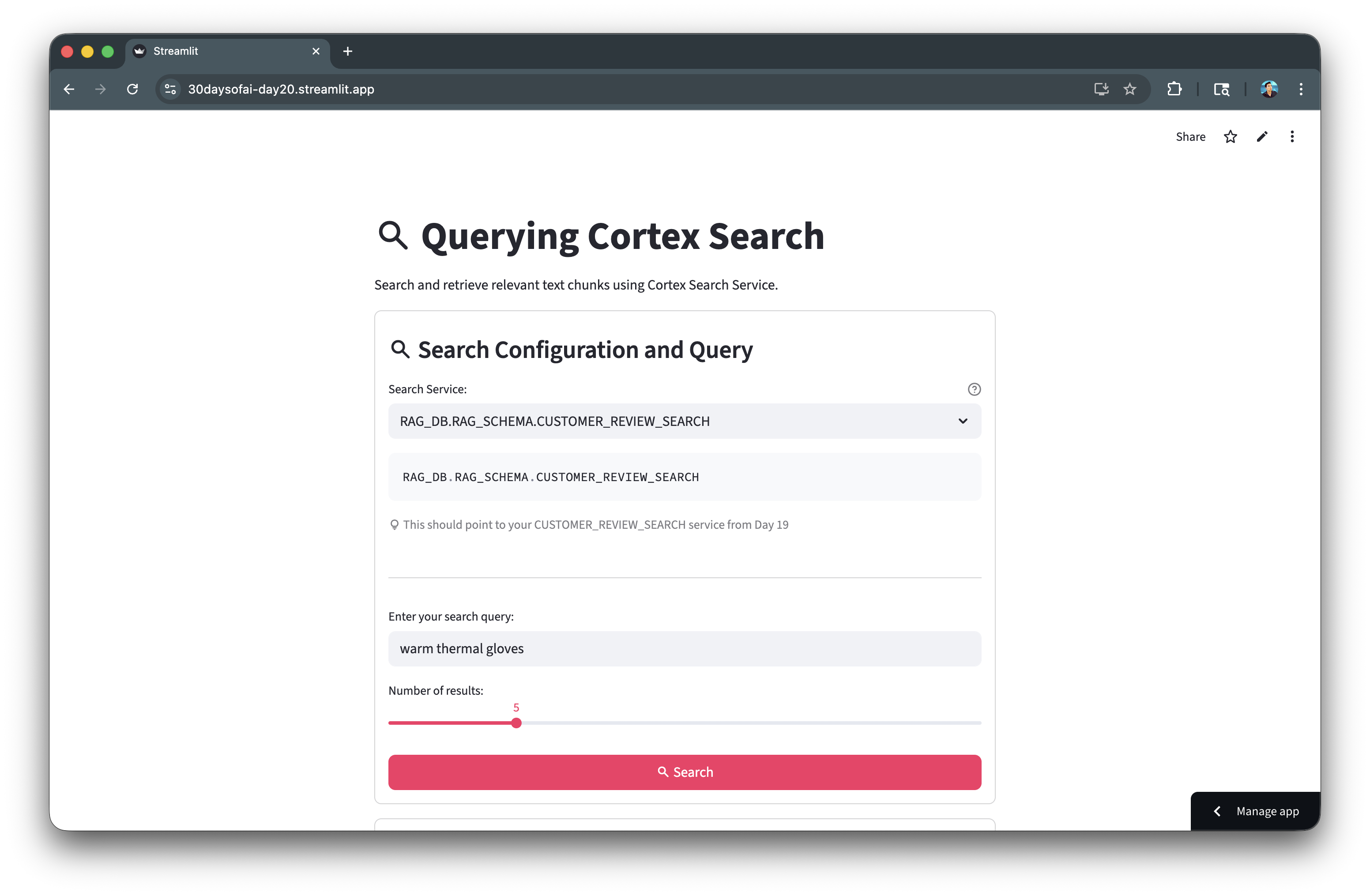
Build RAG Application
Combine retrieval with LLM generation for grounded answers. The RAG pattern retrieves relevant context, then generates responses based on that context.
Note: This section is repurposed from Day 21 of the #30DaysOfAI learning challenge.
RAG Architecture
- Retrieve: Search Cortex Search for relevant chunks
- Augment: Add retrieved chunks to the prompt as context
- Generate: LLM creates an answer grounded in the context
Building the Context
Retrieved chunks are joined with separators to create a context string that's passed to the LLM:
context_chunks = [item.get("CHUNK_TEXT", "") for item in search_results.results] context = "\n\n---\n\n".join(context_chunks)
Prompt Engineering
The prompt explicitly instructs the model to only use provided context, reducing hallucinations:
rag_prompt = f"""You are a helpful assistant. Answer the user's question based ONLY on the provided context. If the context doesn't contain enough information, say "I don't have enough information..." CONTEXT FROM DOCUMENTS: {context} USER QUESTION: {question} """
LLM Completion
Use SNOWFLAKE.CORTEX.COMPLETE() to call the LLM with your context-aware prompt:
response_sql = f""" SELECT SNOWFLAKE.CORTEX.COMPLETE( 'claude-3-5-sonnet', '{rag_prompt.replace("'", "''")}' ) as response """ response = session.sql(response_sql).collect()[0][0]
Full Code
import streamlit as st from snowflake.core import Root try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() def rag_query(question: str, search_service: str, num_chunks: int = 3, model: str = "claude-3-5-sonnet"): root = Root(session) parts = search_service.split(".") svc = root.databases[parts[0]].schemas[parts[1]].cortex_search_services[parts[2]] search_results = svc.search( query=question, columns=["CHUNK_TEXT", "FILE_NAME"], limit=num_chunks ) context_chunks = [item.get("CHUNK_TEXT", "") for item in search_results.results] sources = [item.get("FILE_NAME", "Unknown") for item in search_results.results] context = "\n\n---\n\n".join(context_chunks) rag_prompt = f"""You are a helpful assistant. Answer the user's question based ONLY on the provided context. If the context doesn't contain enough information to answer, say "I don't have enough information to answer that based on the available documents." CONTEXT FROM DOCUMENTS: {context} USER QUESTION: {question} Provide a clear, accurate answer based on the context.""" response_sql = f""" SELECT SNOWFLAKE.CORTEX.COMPLETE( '{model}', '{rag_prompt.replace("'", "''")}' ) as response """ response = session.sql(response_sql).collect()[0][0] return response, context_chunks, sources
Here's the RAG application architecture:
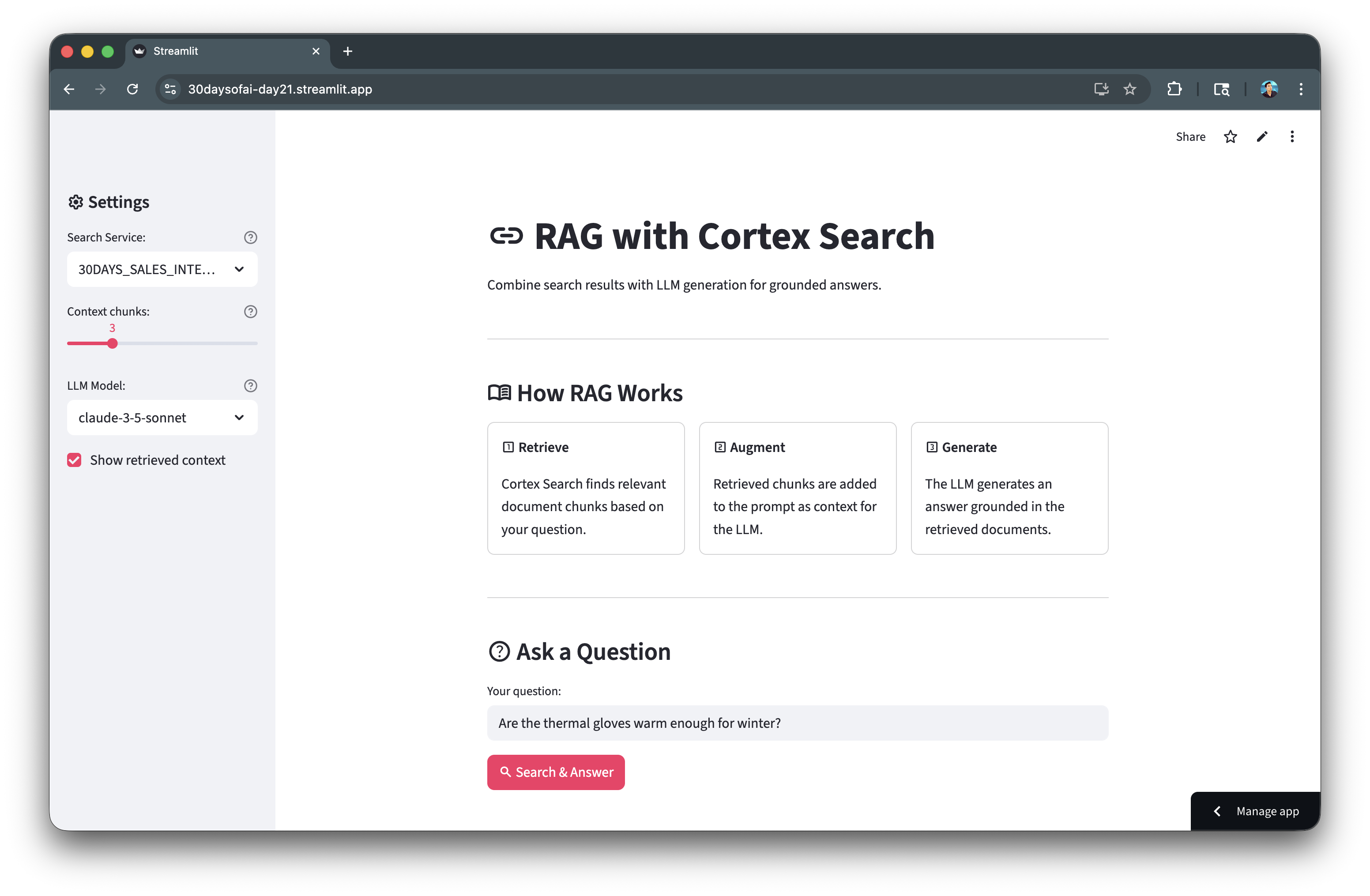
Build RAG Chatbot
Create a conversational interface for your RAG system with chat history and source citations.
Note: This section is repurposed from Day 21 of the #30DaysOfAI learning challenge.
Chat History
Conversation history is stored in session state as a list of message dictionaries. Each message has a role (user/assistant) and content:
if "doc_messages" not in st.session_state: st.session_state.doc_messages = []
Sidebar Configuration
The sidebar provides controls for the search service path, number of context chunks, and a clear chat button:
with st.sidebar: search_service = st.text_input("Cortex Search Service:", value="RAG_DB.RAG_SCHEMA.CUSTOMER_REVIEW_SEARCH") num_chunks = st.slider("Context chunks:", 1, 5, 3)
Source Citations
An expander shows the source chunks used for each answer, providing transparency into retrieval:
with st.expander(f":material/library_books: Sources ({len(chunks)} used)"): for i, (chunk, source) in enumerate(zip(chunks, sources), 1): st.caption(f"**[{i}] {source}**") st.write(chunk[:200] + "...")
Full Code
import streamlit as st from snowflake.core import Root try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() st.title(":material/chat: Chat with Your Documents") if "doc_messages" not in st.session_state: st.session_state.doc_messages = [] with st.sidebar: st.header(":material/settings: Settings") search_service = st.text_input( "Cortex Search Service:", value="RAG_DB.RAG_SCHEMA.CUSTOMER_REVIEW_SEARCH" ) num_chunks = st.slider("Context chunks:", 1, 5, 3) if st.button(":material/delete: Clear Chat"): st.session_state.doc_messages = [] st.rerun() def rag_query(question: str, search_service: str, num_chunks: int = 3): root = Root(session) parts = search_service.split(".") svc = root.databases[parts[0]].schemas[parts[1]].cortex_search_services[parts[2]] results = svc.search(query=question, columns=["CHUNK_TEXT", "FILE_NAME"], limit=num_chunks) context_chunks = [item.get("CHUNK_TEXT", "") for item in results.results] sources = [item.get("FILE_NAME", "Unknown") for item in results.results] context = "\n\n---\n\n".join(context_chunks) rag_prompt = f"""You are a helpful assistant. Answer based ONLY on the provided context. CONTEXT: {context} QUESTION: {question} Answer:""" sql = f"SELECT SNOWFLAKE.CORTEX.COMPLETE('claude-3-5-sonnet', '{rag_prompt.replace(chr(39), chr(39)+chr(39))}')" response = session.sql(sql).collect()[0][0] return response, context_chunks, sources for msg in st.session_state.doc_messages: with st.chat_message(msg["role"]): st.markdown(msg["content"]) if prompt := st.chat_input("Ask about your documents..."): st.session_state.doc_messages.append({"role": "user", "content": prompt}) with st.chat_message("user"): st.markdown(prompt) with st.chat_message("assistant"): with st.spinner("Searching and thinking..."): response, chunks, sources = rag_query(prompt, search_service, num_chunks) st.markdown(response) with st.expander(f":material/library_books: Sources ({len(chunks)} used)"): for i, (chunk, source) in enumerate(zip(chunks, sources), 1): st.caption(f"**[{i}] {source}**") st.write(chunk[:200] + "..." if len(chunk) > 200 else chunk) st.session_state.doc_messages.append({"role": "assistant", "content": response})
Here's the RAG chatbot with sources:

Add Guardrails
Keep your RAG chatbot focused on its domain by adding explicit constraints to the system prompt.
Note: This section is repurposed from Day 22 of the #30DaysOfAI learning challenge.
Why Guardrails?
Without guardrails, users can ask about any topic and the LLM may attempt to answer using its general knowledge. Guardrails keep the assistant focused on the intended domain (customer reviews).
Numbered Guidelines
Explicit numbered instructions tell the LLM exactly what topics to address and how to handle off-topic questions:
STRICT GUIDELINES: 1. ONLY use information from the provided customer review context 2. If asked about unrelated topics, respond: "I can only answer questions about customer reviews..." 3. If context lacks information, say: "I don't have enough information..." 4. Stay focused on: product features, satisfaction, complaints, quality, pricing 5. Do NOT make up information or use outside knowledge
Off-Topic Handling
The guardrails detect off-topic requests and politely redirect the user back to the intended domain.
Full Code
import streamlit as st from snowflake.core import Root try: from snowflake.snowpark.context import get_active_session session = get_active_session() except: from snowflake.snowpark import Session session = Session.builder.configs(st.secrets["connections"]["snowflake"]).create() def rag_query_with_guardrails(question: str, search_service: str, num_chunks: int = 3): root = Root(session) parts = search_service.split(".") svc = root.databases[parts[0]].schemas[parts[1]].cortex_search_services[parts[2]] results = svc.search(query=question, columns=["CHUNK_TEXT", "FILE_NAME"], limit=num_chunks) context = "\n\n---\n\n".join([item.get("CHUNK_TEXT", "") for item in results.results]) rag_prompt = f"""You are a customer review analysis assistant. Your role is to ONLY answer questions about customer reviews and feedback. STRICT GUIDELINES: 1. ONLY use information from the provided customer review context below 2. If asked about topics unrelated to customer reviews (e.g., general knowledge, coding, math, news), respond: "I can only answer questions about customer reviews. Please ask about product feedback, customer experiences, or review insights." 3. If the context doesn't contain relevant information, say: "I don't have enough information in the customer reviews to answer that." 4. Stay focused on: product features, customer satisfaction, complaints, praise, quality, pricing, shipping, or customer service mentioned in reviews 5. Do NOT make up information or use knowledge outside the provided reviews CONTEXT FROM CUSTOMER REVIEWS: {context} USER QUESTION: {question} Provide a clear, helpful answer based ONLY on the customer reviews above.""" sql = f"SELECT SNOWFLAKE.CORTEX.COMPLETE('claude-3-5-sonnet', '{rag_prompt.replace(chr(39), chr(39)+chr(39))}')" response = session.sql(sql).collect()[0][0] return response
Here's the guardrails keeping responses on-topic:
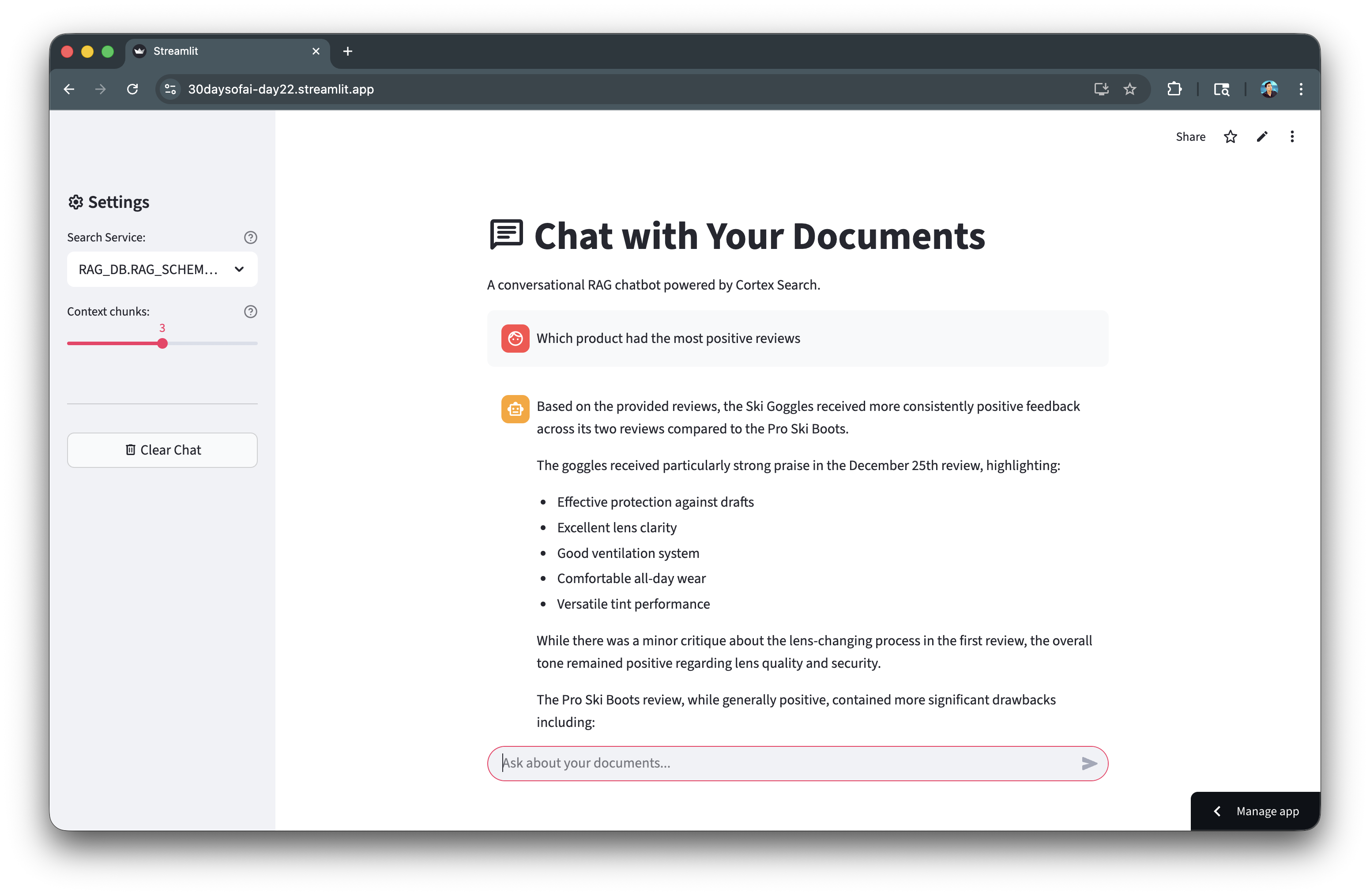
Evaluate with TruLens
TruLens provides automated evaluation of RAG quality using the RAG Triad metrics. This helps you measure and improve retrieval and generation quality.
Note: This section is repurposed from Day 23 of the #30DaysOfAI learning challenge.
RAG Triad Metrics
| Metric | Description |
|---|---|
| Context Relevance | Did we retrieve the right documents? |
| Groundedness | Is the answer based on context (no hallucinations)? |
| Answer Relevance | Does the answer address the question? |
Instrumented RAG Class
The @instrument() decorator marks methods for automatic tracing, capturing inputs, outputs, and timing for each step:
from trulens.core.otel.instrument import instrument class CustomerReviewRAG: @instrument() def retrieve_context(self, query: str) -> str: # Cortex Search retrieval @instrument() def generate_completion(self, query: str, context: str) -> str: # LLM generation @instrument() def query(self, query: str) -> str: # Full RAG pipeline
TruLens Session
Initialize TruLens with a Snowflake connector to store evaluation results in your account:
from trulens.connectors.snowflake import SnowflakeConnector from trulens.core import TruSession tru_connector = SnowflakeConnector(snowpark_session=session) tru_session = TruSession(connector=tru_connector)
Running Evaluations
Register your app and run evaluation against test questions. TruLens automatically computes the RAG Triad metrics:
run.compute_metrics([ "answer_relevance", "context_relevance", "groundedness", ])
Full Code
from trulens.connectors.snowflake import SnowflakeConnector from trulens.core.run import Run, RunConfig from trulens.core import TruSession from trulens.core.otel.instrument import instrument from snowflake.core import Root class CustomerReviewRAG: def __init__(self, snowpark_session): self.session = snowpark_session self.search_service = "RAG_DB.RAG_SCHEMA.CUSTOMER_REVIEW_SEARCH" self.model = "claude-3-5-sonnet" @instrument() def retrieve_context(self, query: str) -> str: root = Root(self.session) parts = self.search_service.split(".") svc = root.databases[parts[0]].schemas[parts[1]].cortex_search_services[parts[2]] results = svc.search(query=query, columns=["CHUNK_TEXT"], limit=3) context = "\n\n".join([r["CHUNK_TEXT"] for r in results.results]) return context @instrument() def generate_completion(self, query: str, context: str) -> str: prompt = f"""Based on this context: {context} Question: {query} Provide a helpful answer:""" prompt_escaped = prompt.replace("'", "''") response = self.session.sql( f"SELECT SNOWFLAKE.CORTEX.COMPLETE('{self.model}', '{prompt_escaped}')" ).collect()[0][0] return response.strip() @instrument() def query(self, query: str) -> str: context = self.retrieve_context(query) answer = self.generate_completion(query, context) return answer tru_connector = SnowflakeConnector(snowpark_session=session) tru_session = TruSession(connector=tru_connector) rag_app = CustomerReviewRAG(session) tru_rag = tru_session.App( rag_app, app_name="customer_review_rag", app_version="v1", main_method=rag_app.query ) test_questions = [ "What do customers say about thermal gloves?", "Are there any durability complaints?", "Which products get the best reviews?" ] run_config = RunConfig( run_name="eval_run_1", dataset_name="test_questions", description="Customer review RAG evaluation" ) run = tru_rag.add_run(run_config=run_config) run.start() run.compute_metrics([ "answer_relevance", "context_relevance", "groundedness", ])
Here's the TruLens evaluation results:
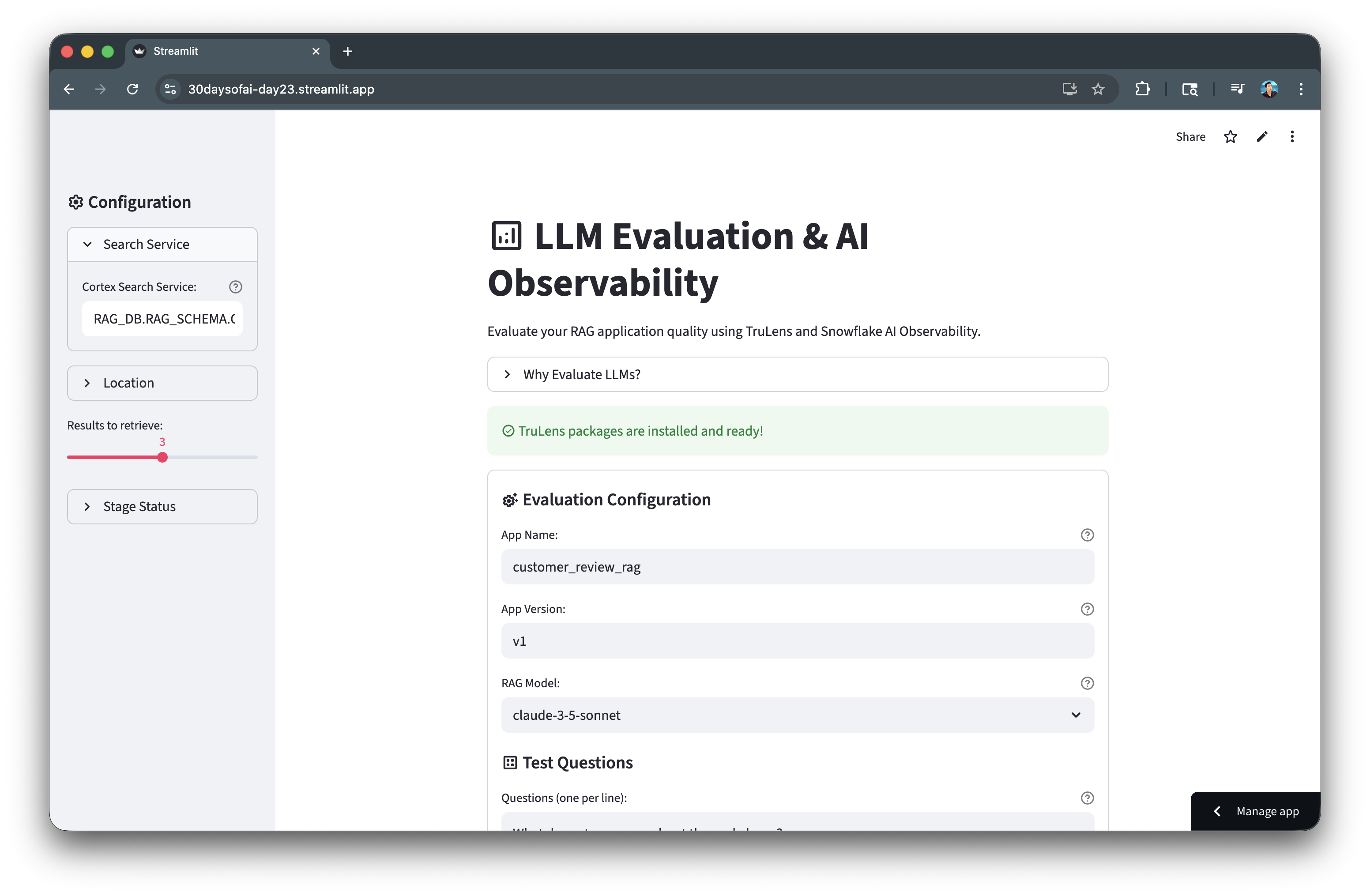
View Results in Snowsight
Navigate to AI & ML → Evaluations → customer_review_rag to see:
- RAG Triad scores per question
- Aggregate metrics across runs
- Trend analysis over time
Deploy the App
Save the complete code above as streamlit_app.py and deploy to your preferred environment:
- Local: Run
streamlit run streamlit_app.pyin your terminal - Streamlit Community Cloud: Deploy your app from a GitHub repository
- Streamlit in Snowflake (SiS): Create a Streamlit app directly in Snowsight
Conclusion And Resources
Congratulations! You've successfully built a complete RAG pipeline with Snowflake Cortex Search. You learned data preparation, chunking strategies, search service creation, building a conversational RAG chatbot with guardrails, and evaluating quality with TruLens.
What You Learned
- Preparing and chunking documents for RAG (Days 16-17)
- Generating embeddings for semantic search (Day 18)
- Creating and configuring Cortex Search services (Day 19)
- Querying Cortex Search for semantic retrieval (Day 20)
- Building RAG applications with grounded responses (Day 21)
- Adding guardrails to keep responses on-topic (Day 22)
- Evaluating RAG quality with TruLens and AI Observability (Day 23)
Related Resources
Documentation:
Additional Reading:
Source Material
This quickstart was adapted from Days 16-23 of the 30 Days of AI challenge:
- Day 16: Document extraction and upload
- Day 17: Data preparation and chunking
- Day 18: Generating embeddings
- Day 19: Creating Cortex Search service
- Day 20: Querying Cortex Search
- Day 21: RAG chatbot
- Day 22: Guardrails for RAG
- Day 23: RAG evaluation with TruLens
Learn more:
This content is provided as is, and is not maintained on an ongoing basis. It may be out of date with current Snowflake instances